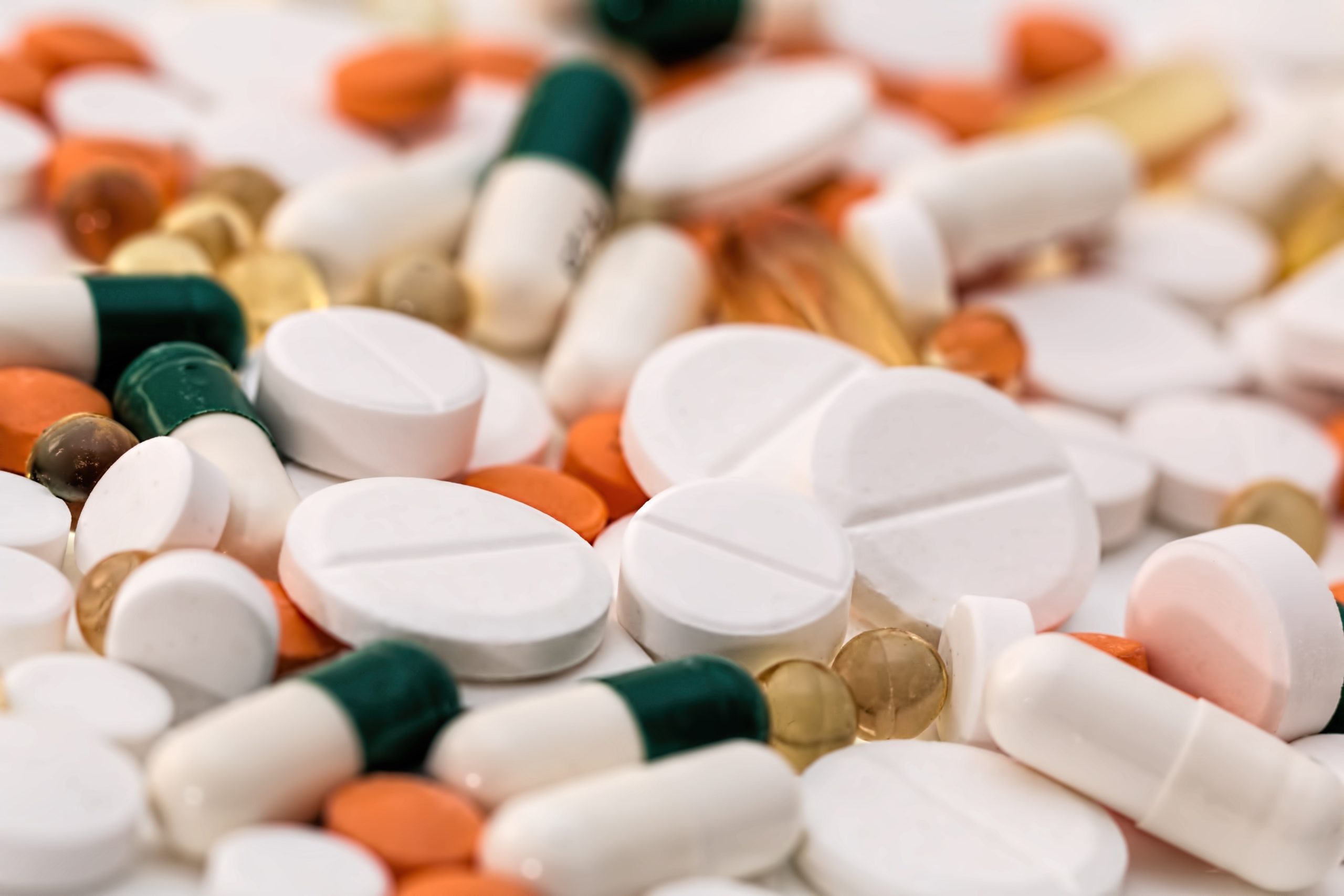
According to Tommi Nyrönen, the correlation between genetics, molecules, and other data of life has only just begun.
“Storing and analysing genomic data at the scale of a million people and connecting this data with other resources – such as data on lifestyle and disease history – will transform our understanding of how diseases are born, how they progress, and most importantly, how they can be cured through individualised treatments,” says Nyrönen.
Nyrönen is Head of the Finnish Node of ELIXIR, the European infrastructure for life science information, and he predicts that the impact of life science data integration will be first seen in the way rare diseases, cancer, and brain-related diseases are diagnosed and treated.
A Scientific e-infrastructure like ELIXIR is essential to the effort of integrating the data of life. It brings together massive life science data sets in Europe, making them securely and reliably accessible for both healthcare and research.
Individualised treatments
Developing individualised treatments is the goal of one of the many projects utilizing the Finnish ELIXIR infrastructure, operated by CSC – IT Centre for Science and connected by the Finnish Research & Education network Funet. Professor Mikko Niemi from the University of Helsinki is developing an algorithm to help predict the necessary dose of adverse effects of a drug, based on patients’ healthcare data combined with their genome information.
But as the genome information is stored in the Biobank of Finland’s National Institute for Health and Welfare and the patient records are stored in the Helsinki University Hospital, just getting access to the data is in itself a complicated undertaking. To create the necessary secure data pathways ELIXIR Finland is collaborating with national partners to create a framework for storing genomics data and interpreting them in clinical practice, for the benefit of Professor Mikko Niemi and other researchers.
Algorithm for more efficiency
It is well known, that people react differently to medications. The reason for that could be the patient’s physical characteristics, his or her genetic makeup, or the influence of other medication. The goal of Professor Mikko Niemi’s research is to develop an algorithm that integrates all these factors and helps doctors find the appropriate drug and the correct dosage for a patient. This would make treatments more effective and reduce side effects.
As an example, Mikko Niemi’s research group is developing decision-making support systems related to pharmacogenetics (Pharmacogenetics is the study of the effect of genes on the efficacy and safety of drug ingredients). The aim is to devise an interpretation algorithm for doctors treating patients with cardiovascular disease to help find the most effective and safe cholesterol medication for each patient. The algorithm uses data on the patient’s characteristics, illnesses, other medications and genome.
Virtual patient
To facilitate the selection of cholesterol medication, Niemi’s research group is building a so-called system pharmacological model.
“It is a kind of virtual patient that can be used to individually predict the effects of each alternative cholesterol drug,” says Mikko Niemi.
“No similar algorithm has been attempted to date. If the algorithm works in the selection of cholesterol medication, a similar way of thinking could also be extended to other drug treatments.”
Access to massive amounts of reliable data is essential for Mikko Niemi and other researchers like him, integrating the data of life to develop more efficient treatment of diseases. The ELIXIR infrastructure helps to reach that goal by giving access to the data, in a standardized way, meeting data protection regulations without compromising openness.
The Finnish ELIXIR node is one of 21 European nodes, altogether involving 650 plus national experts from more than 200 institutes across Europe.
This post is based on a collection of use cases published by ELIXIR Finland. For more information please go to: http://www.elixir-finland.org/en/cases/